
Molecular
mechanisms associated with cutaneous melanoma biology, pathogenesis, and
diagnosis
Josué Mondragón-Morales 1*
1 Escuela Superior de Medicina, Instituto Politécnico
Nacional, Salvador Díaz Mirón esq. Plan de San Luis S/N, Miguel Hidalgo, Casco
de Santo Tomás, 11340, México City, México, CP 11340
Corresponding Authors: Josué
Mondragón-Morales
* Email: jos.mond.m@gmail.com
Abstract
Introduction: Melanoma is considered the most lethal skin cancer, with poor
prognosis in advanced stages. The 2018 World Health Organization (WHO)
Classification classified melanoma into nine different subgroups depending on
the cumulative sun damage, with its respective genetic alterations, which are
necessary to investigate for targeted therapies. Nevertheless, the epigenetic
alterations aren’t included at all in the new molecular classification. It is
understanding the molecular mechanisms associated with melanoma pathogenesis
and its poor prognosis.
Methods: To analyze the molecular mechanisms implicated in melanoma
carcinogenesis, we reviewed the most recent papers using PubMed database and
Google Scholar, the search was carried out using the following medical subject
headings (MeSH) in the search engine: “melanoma
epigenetic mechanisms”, “miRNAs and melanoma”, immunology and melanoma”,
“melanoma pathogenesis”, in combination with boolean
connectors ‘AND’ and ‘OR’. A total of 83 articles were reviewed, published
between 2000 and 2022.
Conclusion: Given the importance of genetic and epigenetic mechanisms implicated
in the prognosis and progression of cancer, this paper aims to review the
literature about its respective regulators, and how they have a relationship
between them in several metabolic, apoptotic, physiological, and biological
processes. It is essential to understand the molecular and immunological
mechanisms involved in melanoma pathogenesis and how the alteration of any of
them leads to the genesis of cancer, to foster the development of novel
targeted therapy strategies.
Keywords: Melanoma, Molecular mechanisms, Skin neoplasm, Genetic, Epigenetic
Introduction
Skin
cancer is the most common form of cancer in the world. It is categorized into
melanoma and non-melanoma skin cancers. Melanoma only accounts for about 1% of
all skin cancers, but is the most aggressive with a poor prognosis, it accounts
90% of all skin cancer deaths, and it’s more frequent in patients between 25
and 40 years old
The
Cancer Genome Atlas (TCGA) classified tumors according to their genomic
characteristics, the most prevalent mutated genes are BRAF, NRAS, NF1-loss and
triple wild type (TWT). However, there are more mutations associated with
tumorigenesis of melanoma, such as CDKN2A (25-35%), TP53 (15%), ARID2 (13.32%),
IDH1, PPP6C, PTEN (14%), DDX3X, RAC1 (9.2%), MAP2K1/2 (10%), RB1, ATRX (9.11%),
SETD2 (9.48%), SF3B1 (33%), TERT (14%), and ERBB2/4 (3.29%)
Modifiable
risk factors
Modifiable
risk factors are related by a high occurrence of oncogenesis, some external
factors such as ultraviolet A (320-400 nm)
Nonmodifiable
risk factors
Genetic
syndromes such as Xeroderma pigmentosum, Neurofibromatosis, Charcot Marie
Tooth, Familiar Atypical Multiple Mole and Melanoma Syndrome (FAMMM) and
Amyotrophic Lateral Sclerosis (ALS)
Genes
involved in melanoma pathogenesis and prognosis
The
mutations of BRAF (incidence of 45%), NRAS (15%), GNAQ, and GNA11 (80-90%)
(involved in the G alpha signaling pathway)
In
addition, ACD, TERF21P, TERF1, TERF2, TINF2 and POT1 are implicated in
telomere maintenance and their mutations increase telomere length and fragility
TP53,
involved in the control of the progression of the cell cycle from G1 to S
phase, its mutation is associated with high risk of melanoma
The
protein phosphatase 2 scaffold subunit A alpha (PPP2R1A) may mediate the
survival and resistance of apoptosis of the type B malignant melanoma cell
lines
microRNAs
(miRNAs) are non-coding RNAs and are important gene regulators. They are
considered as a new potential therapeutic strategy and fundamental prognostic
factor. miR-21-5p reduces cell proliferation and promotes apoptosis by
increasing PDCD4, PTEN, and BTG2. miR-146a-5p is upregulated by BRAF and NRAS,
promoting cell proliferation, cell migration and invasion
Another
molecular mechanism implicated in tumorigenesis of melanoma is the DNA
methylation alterations. The DNA hypermethylation of PTEN, CDKN2A and RASSF1A
have been reported in melanomas. Tellez C.S. et.al reported an elevated
methylation status in their melanoma cell lines: ESR1 (50%), MGMT (50%), RARB2
(44%), RIL (82%), RASSF1A (69%), PAX7 (31%), PGRB (56%), PAX2 (38%), NKX2-3
(63%), OLIG2 (63%), HAND1 (63%), ECAD (88%), CDH13 (44%), and CDKN2A/p16 (6%)
Table 1. The 2018 World Health Organization
(WHO) classification of cutaneous, mucosal, and uveal melanoma
|
Melanomas typically associated with Cumulative
Solar Damage |
Melanomas not consistently associated with
Cumulative Solar Damage |
Nodular melanoma |
|
Pathway I. Superficial spreading melanoma/low-CSD
melanoma |
Pathway IV. Spitz melanoma |
|
|
Pathway II. Lentigo maligna
melanoma/high-CSD melanoma |
Pathway
V. Acral melanoma |
|
|
Pathway III. Desmoplastic melanoma |
Pathway VI. Mucosal melanoma |
|
|
Pathway
VII. Melanomas arising in congenital nevi |
||
|
Pathway VIII. Melanomas arising in blue nevi |
||
|
Pathway
IX. Uveal melanoma |
The
2018 World Health Organization (WHO) classification of cutaneous, mucosal, and
uveal melanoma is based on its arising sun-exposure skin, the role of
ultraviolet (UV) radiation, precursors, and driving and/or recurrent genomic
changes
Pathway
I. Superficial spreading melanoma/low-CSD melanoma
Pathway
I is the route by which melanocytes acquire the genetic aberrations necessary
to become melanoma, however, it is associated with lower CSD. This pathway
contributes to the appearance of superficial spreading melanoma. Superficial
spreading melanoma is the most common form of melanoma. This kind of melanoma
is particularly localized in parts of the body with intermittent sun exposure
like in vacation or weekends. In men, its most frequent localization is in the
back while in women is the back of the legs or calf region. They typically
express BRAF V600E mutations, TERT, and NRAS mutations in less proportion
Pathway
II. Lentigo maligna melanoma/high-CSD melanoma
Pathways
II and III are the pathways necessary to transform melanocytes in melanoma,
however, in contrast with pathway I, these two types of pathways are associated
with high CSD. Through pathway II, melanocytes acquire various genetic
mutations, including NF1, BRAF V600K, NRAS, KIT, CCND1, MITF and TP53 which are
associated with high CSD, and leads to lentigo maligna
melanoma (LMM) transformation. LM is a melanoma subtype considered a melanoma
in situ; it represents about 4-15% of all melanomas. The most frequent site of
this subtype is in head and neck (78.3%). They can be presented as an
amelanotic/hypomelanotic macule or patch, especially in fair-skinned
individuals on chronically sun-damaged skin. There’s described a sex-related
preferential location of LM, developing on the right side of the face in males
and on the left side in females
Pathway
III. Desmoplastic melanoma
As
mentioned above, pathway III is associated with an extremely high mutation
burden with high CSD. Desmoplastic melanoma (DM) arises from this pathway. DM
is a rare variant of cutaneous melanoma; it accounts for about 1% of all
melanomas. They’re commonly amelanotic or sparsely pigmented and are typically
endophytic
Pathway
IV. Spitz melanoma
Previously
to WHO classification, Spitz melanoma (SM) was classified based on the
cytomorphologic features in spitzoid melanomas.
Nowadays, SMs are classified based on their morphologic and genomic alterations
such as HRAS, ALK, NTRK1, MAP3K8, BRAF, and CDKN2A mutations, in contrast with
its counterpart Spitz Nevi (SN). SMs are rare, they represent about 1-2% of all
melanocytic lesions. The mean age of diagnosis in SM is 22 years old. They can
be localized in any part of the body but is more frequent in lower extremities
(40-50%), trunk (20%), upper limbs (15%), and head/neck (5%). SM are elevated
lesions, mostly of them are larger than 1 cm in diameter and can have pink to
black coloration. The majority are asymmetrical, with coloration variety,
present shiny white lines, and polymorphous vascular patterns.
Pathway
V. Acral melanoma
Acral
melanomas arise on the non-hair bearing skin, especially in the lower
extremities (78%), comprises about 2-3% of all melanomas. They have a high
number of structural chromosomal changes and lower frequencies of BRAF
mutations (10-23%), KIT mutations (3-29%), amplification of CCND1 and CDK4, and
deletion/mutations in CDKN2A, PTEN, NF1 and hTERT
Pathway
VI. Mucosal melanoma
Primary
mucosal melanomas (MM) are derived from neural crest cells that migrate to
several sites, they can be found in the respiratory, gastrointestinal, and
genitourinary tract, it represent about 0.8-3.7% of
all melanomas. They are associated with aggressive and less favorable prognoses.
C-KIT is overexpressed (80%), BRAF mutations are less common (<10%) and
SF3B1 mutations (12%) cause directly aberrant gene transcripts which lead to
mRNA degradation or abnormal protein function in MM. There are some specific
risk factors such as tobacco, and formaldehyde (associated with oral and sinonasal MM), and human immunodeficiency virus (HIV)
infection (associated with rectal MM)
Pathway
VII. Melanoma arising from congenital melanocytic nevi
Congenital
melanocytic nevi (CMN) are hamartomas of the neuroectoderm, they are seen in
about 1-6% of all birth, and they are caused by genetic mosaicism. Large/giant
CMN occur in 1/20,000-50,000 births. They can be classified by its size in
small (<1.5 cm), medium (1.5-20 cm), and large (>20 cm). BRAF mutations
are mostly presented in small nevi, and NRAS mutations in large/giant CMN.
Melanoma risk is difficult to quantify, but there is a high risk in lesions
that lie across the spine or those who has numerous satellite lesions (10-15%
of risk)
Pathway
VIII. Melanoma arising from blue nevi
As
mentioned, pathway VIII is an UV-unrelated group. This type of pathway is
associated with chromosomal aberrations added to a precursor lesion, blue nevi.
Blue Nevis are uncommon lesions. They express GNAQ and GNA11 mutations, and
infrequently in PLCB4 or CYSLTR2, EIF1AX, SF3B1 and BAP1 mutations. In
addition, the gain of chromosomal arms 1q, 4p, 6p and losses of 1p and 4q have
been identified
Pathway
IX. Uveal melanoma
The
eye is an immune-privileged organ, so, intraocular environment is considered an
immunosuppressive environment, where melanoma can proliferate, invade, and
progress to metastasis. Uveal melanoma (UM) is a rare disease, and it has been
demonstrated that it is different from its cutaneous counterpart. More than 90%
involve choroid, 6% are confined to the ciliary body and 4% to the iris. They
represent the most frequent intraocular primary tumor in adults
Pathway
X. Nodular melanoma
Nodular
melanomas arise from any of the pathways mentioned above, that’s why they have
heterogeneous epidemiologic and genomic features. They are characterized to be
nodular or papular at the clinical examination, with
homogeneous or heterogeneous pigment. BRAF and NRAS mutations have been
demonstrated in these kinds of tumors, however, its genomic alterations are
still unknown
Nowadays,
there is a molecular classification of melanoma, with prognostic importance,
however it has not yet been added to the current WHO classification.
I.
BRAF-mutant: about 60% presents
CDKN2A mutation, TP53 mutation (10%), ARID2 mutated (15%), PPP6C mutated (10%),
PDL1 and MITF amplification
II.
RAS-mutant: CDKN2A mutated (about
70%), CCDN1 amplification (10%), TP53 mutation (20%), ARID2 mutation (15%) and
PPP6C mutation (15%)
III.
NF1-mutant: CDKN2A mutation (70%),
RB1 mutation (10%), TP53 mutation (30%), and ARID2 mutation (30%)
IV.
Triple Wildtype: CDKN2A mutation
(40%), CDK4 amplification (15%), CCDN1 amplification (10%), and MDM2
amplification (15%)
Tumorigenesis
in melanoma cells is regulated by multiple signaling pathways, modulated by
genetic and epigenetic mechanisms, with a straight interrelation between them

Figure 1. Genetic and
epigenetic mechanisms of malignancy in melanoma.
Molecular mechanisms implicated in pathogenesis
Melanoma
biology
Melanocytes
are a heterogeneous group of cells, derived from the neural crest. They produce
the protective skin-darkening pigment melanin in epidermis, hair, and iris,
which is responsible of the protection of DNA from UV-mediated damage
Cutaneous
melanoma is the most aggressive skin cancer; it derives from melanocytes. It
accounts about 90% of melanomas including mucosal and uveal melanomas and
represents about 1% of all skin cancers. The biology of the tumor is associated
with the microenvironment, it has been demonstrated the hypoxic and acidity of
microenvironment as an important role in melanoma biology
Acidosis
plays an additional role by the dedifferentiation of cancer cells, to an
immature phenotype, commonly known as Cancer Stem-like Cells (CSC), with the
ability to self-renew and keep them in a quiescent state responsible for
chemotherapy and radiotherapy resistance
Nowadays,
it is demonstrated that lipid metabolism is implicated in promoting melanoma
progression. Carnitine palmitoyltransferase 2 (CPT2),
phospholipase D3 (PLD3), inositol triphosphate protein kinase B (ITPKB), and
inositol triphosphate receptor 3 (ITRP3), genes that encode lipid metabolism
proteins, are significantly upregulated genes in melanomas compared with benign
nevi, and their expression is associated with melanoma pathogenesis. However,
the role of this kind of proteins in melanoma pathogenesis is still unclear
Microenvironment
Melanoma
is one of the most immunogenic tumors, so its microenvironment has a high
concentration of infiltrating immune cells, however, most of them, are
inhibitory immune populations, including T regulatory (T reg) cells,
tissue-associated macrophages (TAMs) and myeloid-derived immunosuppressive
cells (MDSCs)
The
most frequent inflammatory cells in the melanoma microenvironment are CD163+
histiocytes, CD3+ T lymphocytes, CD68+ histiocytes, cytotoxic CD8+ T
lymphocytes, CD4+ regulatory T cells
The
immune system has an efficient recognition of tumor cells, by presenting
melanoma antigens to T cells, which can expand and become effector melanoma-specific
T cells. Two immune checkpoints can upregulate or downregulate the immune
stimulation: cytotoxic T lymphocyte antigen 4 (CTLA-4), a coinhibitory molecule
on T cells that inhibits cells activation by ligation with CD86 and CD80; and
programmed death 1 (PD-1), another immune checkpoint, that can be inhibited by
programmed death 1 ligand (PD-L1 and PD-L2) expressed in tumor cells
In
addition to PD-1, CTLA-4 is the second most frequently known immune suppressive
checkpoint regulator, its function is associated with immune suppressive
activities by inhibiting T cell activation. CTLA-4 outcompetes CD28 for the
ligands, CD80/CD86, in consequence, T cells become anergic
Diagnosis
The
diagnostic approach starts with dermoscopic
evaluation, it’s necessary to describe the skin lesion with the mnemotechnic
ABCDE (Asymmetry (the most common criterion: 84.5%), Border, Color (the
multicomponent pattern is the most characteristic and most common patient
associated with melanoma), Diameter and Evolution) as seen in Figure 2. Dermoscopy is a fundamental for early diagnosis and in the
preoperative estimate of the Breslow index
Most
patients with cutaneous melanoma are asymptomatic, and they come to clinical
care only in the presence of a suspicious injury. At the same time, patients
with UM are asymptomatic (>40%), those who present symptoms may develop
blurred or distorted vision, visual field loss of photopsia, or other ocular
symptoms, rarely large tumors induce vitreous hemorrhage
The
visual inspection of a suspicious skin lesion is the first step in melanoma
diagnosis, its sensitivity is about 76% (66-85%) and specificity 75%
(57-87%)
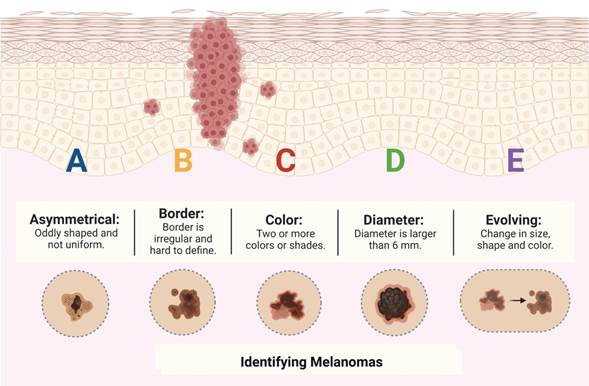
Figure 2. ABCDE for
identifying melanoma.
Table 2. Accuracy of several methods used
in melanoma diagnosis and staging.
|
Diagnostic method. |
Accuracy. |
Characteristics. |
|
|
Sensitivity. |
Specificity. |
||
|
Visual inspection |
76% (66-85%) |
75% (57-87%) |
Clinical inspection of pigmented skin
lesions using the mnemonic ABCDE |
|
Dermoscopy |
Without Artificial Intelligence Support
(53.3-65.5%) With Artificial Intelligence Support
(81.9-87.6%) |
Without Artificial Intelligence Support
(62.3-78.9%) With Artificial Intelligence Support
(74.8-83.4%) |
It’s the examination of pigmented and
non-pigmented skin lesions with the naked eye With artificial intelligence support like
reflectance confocal microscopy increases accuracy |
|
Histopathology |
91% (84-95%) |
94% (86-98%) |
The histological examination of a pigmented
skin lesion. It’s considered the gold standard for melanoma diagnosis |
|
Immunohistochemistry (IHC) |
Adjuvant to histopathology, it consists in
the examination of melanoma antigens using anti-H4K20me and anti-H3K27me3
monoclonal antibodies, which interact with their respective antigens |
||
|
Comparative Genomic Hybridization (CGH) |
92-96% |
87-100% |
Adjuvant to histopathology detects
genome-wide changes in DNA copy number, but it doesn’t detect actual
mutations. It can detect genetic anomalies in chromosomes 6p, 1q, 7p, 7q, 8q,
17q and 20q and/or losses of 9p, 9q, 10q, 10p, 6q and 11q |
|
Fluorescent In Situ Hybridization (FISH) |
43-100% |
29-80% |
Adjuvant to histopathology detects
cytogenetic abnormalities by direct visualization |
|
Ultrasound. (US) |
Nodal metastasis 35.4% (17-59.4%) |
Nodal metastasis 93.9% (86.1-97.5%) |
Ultrasound uses high-frequency sound waves
to create images in the body, it can be used to assist in detection of lymph
node metastasis |
|
Ultrasound with Fine Needle Aspiration
Cytology (US FNAC) |
Nodal metastasis 18% (3.58-56.5%) |
Nodal metastasis 99.8% (99.1-99.9%) |
The cytologic examination of skin lesions
using a fine needle aspiration guided by ultrasound |
|
Computed Tomography (CT) |
Nodal metastasis 87.2 (76.5-93.4%). Distant metastasis 73.4% (63.6-81.3%) |
Nodal metastasis 69.2% (34.6-90.5%). Distant metastasis 72% (64.3-78.5%) |
Uses ionizing radiation in the form of
X-rays to take cross sectional images of the body, is used to evaluate
metastasis |
|
Magnetic Resonance Imaging. (MRI) |
Nodal metastasis 83.7% (68.8-92.3%). Distant metastasis 74.5% (62.1-83.9%) |
Nodal metastasis 77.7% (72.4-82.1%). Distant metastasis 85.8% (70.4-93.9%) |
Uses large magnets and non-ionizing
radiation in the form of radio waves to generate images of the body, is used
to evaluate metastasis |
|
Positron Emission Tomography (PET/CT). |
Nodal metastasis 86.5% (80.2-91.1%). Distant metastasis 92.5% (85.3-96.4%). Detection of bone metastasis 90.2%
(78.5-95.9%) |
Nodal metastasis 92.5% (85-96.4%). Distant metastasis 89.7% (78.8-95.3%). Detection of bone metastasis 88.2%
(72.5-95.5%) |
A nuclear medicine imaging technique, it
uses a radioactive component (18FDG intravenous) which is taken up
by cancer cells |
New
treatment strategies
Most
patients are diagnosed in early-stage disease, in which surgical excision is
the treatment of choice, because it’s curative in most of the cases
BRAF
is a serine/threonine protein kinase, encoded on chromosome 7q34, which
activates the MAPK/ERK-signaling pathway. The most frequent BRAF mutation (90%)
is located at codon 600, in which a single nucleotide mutation results in the substitution
of glutamic acid for valine (V600E)
As
mentioned above, melanoma cells express PD-L1 in their membrane surfaces, and
the interaction of CTLA-4 in T cells membrane surfaces results in T cell
anergy. These two immune checkpoints are important for an effective immune
response. Immune checkpoint inhibitors play key roles, when a tumor does not
have targeted mutations, or it does not respond to BRAF/MEK inhibitors. There
are two types of immune checkpoint inhibitors, PD-1 inhibitors (nivolumab and
pembrolizumab) and CTLA-4 antibody inhibitors (ipilimumab). The inhibition of
these two immune checkpoints helps the immune system to recognize cancer cells
by suppressing melanoma's immune evasion system
Future
directions
Numerous phase
I and II clinical trials are currently underway to explore innovative agents
and multimodal approaches to enhance the prognosis of patients facing melanoma.
Many of these trials are centered on monoclonal antibodies, which represent
vital components of targeted strategies in the era of precision medicine. While
monoclonal antibodies hold considerable promise, their mechanism of action
often entails inhibiting critical pathways associated with melanoma
pathogenesis. Consequently, these interactions can lead to adverse effects.
Discussion
Various
researchers have conducted exhaustive investigations into the mechanisms
discussed earlier, underscoring their significance in driving carcinogenesis in
melanocytes and their correlation with various molecular subclassifications.
While new treatment strategies have emerged based on these mechanisms, some
still lack targeted therapies, necessitating further research into the yet
uncharted direct and indirect contributors to tumorigenesis. Genetic,
epigenetic alterations and tumor microenvironment have all been associated with
this unfavorable prognosis due to their facilitation of uncontrolled
proliferation of malignant cells. Therefore, this article seeks to consolidate
valuable insights on melanoma, to contribute to the formulation of treatment strategies.
Conclusions
Melanoma
is the most aggressive skin cancer, with poor prognosis and high mortality. Its
pathogenesis encompasses many molecular mechanisms, incorporating genetic and
epigenetic factors. These mechanisms operate within various signaling pathways,
often displaying interconnectedness and interplay. They exert their influence
on pro- and anti-apoptotic proteins, sculpting the microenvironment by
regulating cell proliferation, invasiveness, and immune evasion. Intriguingly,
these emerging mechanisms are not confined to melanoma but are also observed in
other solid tumors, including breast, colorectal, urogenital, pancreatic, and
lung tumors. Nowadays, these new molecular mechanisms open the possibility of
investigating new alternatives for possible targeted therapies. The primary
objective of this review article is to provide a comprehensive account of the
molecular mechanisms involved in melanoma pathogenesis and how the alteration
of any of them leads to the genesis of cancer, to foster the development of
novel targeted therapy strategies.
Author contribution
This
manuscript was written entirely by JMM.
Conflict of interest
The
authors report no conflict of interest.
References
1. Dubbini N, Puddu
A, Salimbeni G, Malloggi S,
Gandini D, Massei P, et al. Melanoma prevention: Comparison of different
screening methods for the selection of a high risk
population. Int J Environ Res Public Health. 2021 Feb 2;18(4):1–10.
2. Eddy K, Chen S. Overcoming immune evasion
in melanoma. Vol. 21, International Journal of Molecular Sciences. MDPI AG;
2020. p. 1–48.
3. Oba J, Woodman SE. The genetic and
epigenetic basis of distinct melanoma types. Vol. 48, Journal of Dermatology.
Blackwell Publishing Ltd; 2021. p. 925–39.
4. Amalinei C, Grigoraș A, Lozneanu L, Căruntu ID, Giușcă SE, Balan RA.
The Interplay between Tumour Microenvironment
Components in Malignant Melanoma. Vol. 58, Medicina
(Lithuania). MDPI; 2022.
5. Emri G, Paragh G,
Tósaki Á, Janka E, Kollár S, Hegedűs C, et al.
Ultraviolet radiation-mediated development of cutaneous melanoma: An update. J Photochem Photobiol B. 2018 Aug 1;185:169–75.
6. Dzwierzynski WW.
Melanoma Risk Factors and Prevention. Vol. 48, Clinics in Plastic Surgery. W.B.
Saunders; 2021. p. 543–50.
7. Raimondi S, Suppa M, Gandini S. Melanoma
epidemiology and sun exposure. Acta Derm Venereol. 2020;100(100-year theme Skin malignancies):250–8.
8. Bataille V. It’s not all sunshine: Non-sun-related melanoma risk-factors. Vol. 100, Acta
Dermato-Venereologica. Medical Journals/Acta D-V;
2020. p. 259–65.
9. Warner AB, McQuade JL. Modifiable Host
Factors in Melanoma: Emerging Evidence for Obesity, Diet, Exercise, and the
Microbiome. Vol. 21, Current Oncology Reports. Current Medicine Group LLC 1;
2019.
10. O’Neill CH, Scoggins CR. Melanoma. Vol. 120,
Journal of Surgical Oncology. John Wiley and Sons Inc.; 2019. p. 873–81.
11. Carr S, Smith C, Wernberg J. Epidemiology and
Risk Factors of Melanoma. Vol. 100, Surgical Clinics of North America. W.B.
Saunders; 2020. p. 1–12.
12. Motofei IG.
Malignant Melanoma: Autoimmunity and Supracellular Messaging as New Therapeutic
Approaches. Vol. 20, Current Treatment Options in Oncology. Springer New York
LLC; 2019.
13. van Poppelen NM, de Bruyn DP, Bicer T, Verdijk R, Naus N, Mensink H, et al. Genetics of ocular
melanoma: Insights into genetics, inheritance and testing. Vol. 22,
International Journal of Molecular Sciences. MDPI AG; 2021. p. 1–19.
14. Pawlowska E, Szczepanska
J, Szatkowska M, Blasiak J. An interplay between senescence, apoptosis and
autophagy in glioblastoma multiforme—role in pathogenesis and therapeutic
perspective. Vol. 19, International Journal of Molecular Sciences. MDPI AG;
2018.
15. Quan VL, Panah E, Zhang B, Shi K, Mohan LS,
Gerami P. The role of gene fusions in melanocytic neoplasms. Vol. 46, Journal
of Cutaneous Pathology. Blackwell Publishing Ltd; 2019. p. 878–87.
16. Toussi A, Mans N,
Welborn J, Kiuru M. Germline mutations predisposing to melanoma. Vol. 47,
Journal of Cutaneous Pathology. Blackwell Publishing Ltd; 2020. p. 606–16.
17. Fang S, Lu J, Zhou X, Wang Y, Ross MI, Gershenwald JE, et al. Functional annotation of melanoma
risk loci identifies novel susceptibility genes. Carcinogenesis. 2020 Jun
17;41(4):452–7.
18. Chen J, Sun W, Mo N, Chen X, Yang L, Tu S, et
al. Identification of key genes involved in the pathogenesis of cutaneous
melanoma using bioinformatics analysis. Journal of International Medical
Research. 2020 Jan 1;48(1).
19. Kulesza DW, Ramji K, Maleszewska
M, Mieczkowski J, Dabrowski M, Chouaib S, et al.
Search for novel STAT3-dependent genes reveals SERPINA3 as a new STAT3 target
that regulates invasion of human melanoma cells. Laboratory Investigation. 2019
Nov 1;99(11):1607–21.
20. Teixido C, Castillo P, Martinez-Vila C, Arance A, Alos L. Molecular
markers and targets in melanoma. Vol. 10, Cells. MDPI; 2021.
21. Dika E, Riefolo M,
Porcellini E, Broseghini E, Ribero S, Senetta R, et
al. Defining the Prognostic Role of MicroRNAs in Cutaneous Melanoma. Journal of
Investigative Dermatology. 2020 Nov 1;140(11):2260–7.
22. Santourlidis S,
Schulz WA, Araúzo-Bravo MJ, Gerovska
D, Ott P, Bendhack ML, et al. Epigenetics in the
Diagnosis and Therapy of Malignant Melanoma. Vol. 23, International Journal of
Molecular Sciences. MDPI; 2022.
23. Elder DE, Bastian BC, Cree IA, Massi D, Scolyer RA. The 2018 World Health Organization
classification of cutaneous, mucosal, and uveal melanoma detailed analysis of 9
distinct subtypes defined by their evolutionary pathway. Vol. 144, Archives of
Pathology and Laboratory Medicine. College of American Pathologists; 2020. p.
500–22.
24. Deluca E V., Deluca E V., Perino F, Distefani A, Coco V, Fossati B, et al. Lentigo maligna: Diagnosis and treatment. Vol. 155, Giornale Italiano di Dermatologia
e Venereologia. Edizioni
Minerva Medica; 2020. p. 179–89.
25. Iznardo H, Garcia-Melendo C, Yélamos O. Lentigo maligna: Clinical presentation and appropriate management.
Vol. 13, Clinical, Cosmetic and Investigational Dermatology. Dove Medical Press
Ltd; 2020. p. 837–55.
26. Cheng TW, Ahern MC, Giubellino
A. The Spectrum of Spitz Melanocytic Lesions: From Morphologic Diagnosis to
Molecular Classification. Front Oncol. 2022 Jun 7;12.
27. Chen YA, Teer JK, Eroglu Z, Wu JY, Koomen JM,
Karreth FA, et al. Translational pathology, genomics
and the development of systemic therapies for acral melanoma. Vol. 61, Seminars
in Cancer Biology. Academic Press; 2020. p. 149–57.
28. Yde SS, Sjoegren P,
Heje M, Stolle LB. Mucosal Melanoma: a Literature
Review. Vol. 20, Current Oncology Reports. Current Medicine Group LLC 1; 2018.
29. Ma Y, Xia R, Ma X, Judson-Torres RL, Zeng H.
Mucosal Melanoma: Pathological Evolution, Pathway Dependency and Targeted
Therapy. Vol. 11, Frontiers in Oncology. Frontiers Media S.A.; 2021.
30. Farabi B, Akay BN, Goldust M, Wollina U, Atak MF,
Rao B. Congenital melanocytic naevi: An up-to-date overview. Vol. 62,
Australasian Journal of Dermatology. Blackwell Publishing; 2021. p. e178–91.
31. Moustafa D, Blundell AR, Hawryluk EB.
Congenital melanocytic nevi. Vol. 32, Current opinion in pediatrics. NLM
(Medline); 2020. p. 491–7.
32. Ferrara G, Argenziano G. The WHO 2018
Classification of Cutaneous Melanocytic Neoplasms: Suggestions From Routine Practice. Vol. 11, Frontiers in Oncology.
Frontiers Media S.A.; 2021.
33. Elder DE, Bastian BC, Cree IA, Massi D, Scolyer RA. The 2018 World Health Organization
classification of cutaneous, mucosal, and uveal melanoma detailed analysis of 9
distinct subtypes defined by their evolutionary pathway. Vol. 144, Archives of
Pathology and Laboratory Medicine. College of American Pathologists; 2020. p.
500–22.
34. Branisteanu D, Bogdanici
C, Branisteanu D, Maranduca M, Zemba M, Balta F, et
al. Uveal melanoma diagnosis and current treatment options (Review). Exp Ther
Med. 2021 Oct 11;22(6).
35. Jager MJ, Shields CL, Cebulla CM,
Abdel-Rahman MH, Grossniklaus HE, Stern MH, et al.
Uveal melanoma. Nat Rev Dis Primers. 2020 Dec 1;6(1).
36. Tímár J, Ladányi A. Molecular Pathology of
Skin Melanoma: Epidemiology, Differential Diagnostics, Prognosis and Therapy
Prediction. Vol. 23, International Journal of Molecular Sciences. MDPI; 2022.
37. Boussadia Z,
Gambardella AR, Mattei F, Parolini I. Acidic and hypoxic microenvironment in
melanoma: Impact of tumour exosomes on disease
progression. Vol. 10, Cells. MDPI; 2021.
38. Napoli S, Scuderi C, Gattuso G, Bella V Di,
Candido S, Basile MS, et al. Functional Roles of Matrix Metalloproteinases and
Their Inhibitors in Melanoma. Vol. 9, Cells. NLM (Medline); 2020.
39. Ostrowski SM, Fisher DE. Biology of Melanoma.
Vol. 35, Hematology/Oncology Clinics of North America. W.B. Saunders; 2021. p.
29–56.
40. Brito FC, Kos L. Timeline and distribution of
melanocyte precursors in the mouse heart. Pigment Cell Melanoma Res. 2008
Aug;21(4):464–70.
41. Fischer GM, Vashisht Gopal YN, McQuade JL,
Peng W, DeBerardinis RJ, Davies MA. Metabolic strategies of melanoma cells:
Mechanisms, interactions with the tumor microenvironment, and therapeutic
implications. Vol. 31, Pigment Cell and Melanoma Research. Blackwell Publishing
Ltd; 2018. p. 11–30.
42. Ekström EJ, Bergenfelz
C, von Bülow V, Serifler F, Carlemalm
E, Jönsson G, et al. WNT5A induces release of exosomes containing
pro-angiogenic and immunosuppressive factors from malignant melanoma cells. Mol
Cancer. 2014 Apr 26;13(1).
43. Hood JL. Melanoma exosome induction of
endothelial cell GM-CSF in pre-metastatic lymph nodes may result in different
M1 and M2 macrophage mediated angiogenic processes. Med Hypotheses. 2016 Sep 1;94:118–22.
44. Li J, Chen J, Wang S, Li P, Zheng C, Zhou X,
et al. Blockage of transferred exosome-shuttled miR-494 inhibits melanoma
growth and metastasis. J Cell Physiol. 2019 Sep 1;234(9):15763–74.
45. Alegre E, Sanmamed
MF, Rodriguez C, Carranza O, Martín-Algarra S, González Á. Study of circulating
MicroRNA-125b levels in serum exosomes in advanced melanoma. Arch Pathol Lab Med. 2014;138(6):828–32.
46. Xiao D, Barry S, Kmetz D, Egger M, Pan J, Rai
SN, et al. Melanoma cell-derived exosomes promote epithelial-mesenchymal
transition in primary melanocytes through paracrine/autocrine signaling in the
tumor microenvironment. Cancer Lett. 2016 Jul 1;376(2):318–27.
47. Zhou X, Yan T, Huang C, Xu Z, Wang L, Jiang
E, et al. Melanoma cell-secreted exosomal miR-155-5p
induce proangiogenic switch of cancer-associated fibroblasts via
SOCS1/JAK2/STAT3 signaling pathway. Journal of Experimental and Clinical Cancer
Research. 2018 Oct 3;37(1).
48. Marzagalli M,
Raimondi M, Fontana F, Montagnani Marelli M, Moretti RM, Limonta P. Cellular
and molecular biology of cancer stem cells in melanoma: Possible therapeutic
implications. Vol. 59, Seminars in Cancer Biology. Academic Press; 2019. p.
221–35.
49. Hanahan D. Hallmarks of Cancer: New
Dimensions. Cancer Discov [Internet]. 2022 Jan 1
[cited 2023 Oct 23];12(1):31–46. Available from:
https://dx.doi.org/10.1158/2159-8290.CD-21-1059
50. Cheng N, Chytil A, Shyr Y, Joly A, Moses HL.
Transforming growth factor-beta signaling-deficient fibroblasts enhance
hepatocyte growth factor signaling in mammary carcinoma cells to promote
scattering and invasion. Mol Cancer Res. 2008 Oct 1;6(10):1521–33.
51. Tan SH, Barker N. Stemming Colorectal Cancer
Growth and Metastasis: HOXA5 Forces Cancer Stem Cells to Differentiate. Cancer
Cell. 2015 Dec 14;28(6):683–5.
52. Perekatt AO, Shah
PP, Cheung S, Jariwala N, Wu A, Gandhi V, et al. SMAD4 Suppresses WNT-Driven
Dedifferentiation and Oncogenesis in the Differentiated Gut Epithelium. Cancer
Res. 2018 Sep 1;78(17):4878–90.
53. Yu X, Qiu W, Yang L, Zhang Y, He M, Li L, et
al. Defining multistep cell fate decision pathways during pancreatic
development at single-cell resolution. EMBO J. 2019 Apr 15;38(8).
54. Darwiche N. Epigenetic mechanisms and the
hallmarks of cancer: an intimate affair. Am J Cancer Res. 2020;10(7):1954.
55. Wang B, Kohli J, Demaria M. Senescent Cells
in Cancer Therapy: Friends or Foes? Trends Cancer. 2020 Oct 1;6(10):838–57.
56. Tao J, Li Y, Liu YQ, Wang L, Yang J, Dong J,
et al. Restoration of the expression of transports associated with antigen
processing in human malignant melanoma increases tumor-specific immunity.
Journal of Investigative Dermatology. 2008;128(8):1991–6.
57. Gajewski TF, Schreiber H, Fu YX. Innate and
adaptive immune cells in the tumor microenvironment. Vol. 14, Nature
Immunology. 2013. p. 1014–22.
58. Fujii H, Josse J, Tanioka M, Miyachi Y,
Husson F, Ono M. Regulatory T Cells in Melanoma Revisited by a Computational
Clustering of FOXP3+ T Cell Subpopulations. The Journal of Immunology. 2016 Mar
15;196(6):2885–92.
59. Magnuson AM, Kiner E, Ergun A, Park JS, Asinovski N, Ortiz-Lopez A, et al. Identification and
validation of a tumor-infiltrating Treg transcriptional signature conserved
across species and tumor types. Proc Natl Acad Sci U
S A. 2018 Nov 6;115(45):E10672–81.
60. Viguier M, Lemaître
F, Verola O, Cho MS, Gorochov
G, Dubertret L, et al. Foxp3 Expressing CD4+CD25high
Regulatory T Cells Are Overrepresented in Human Metastatic Melanoma Lymph Nodes
and Inhibit the Function of Infiltrating T Cells. The Journal of Immunology.
2004 Jul 15;173(2):1444–53.
61. Umansky V, Sevko A,
Gebhardt C, Utikal J. Myeloid-derived suppressor
cells in malignant melanoma. Vol. 12, JDDG - Journal of the German Society of
Dermatology. Wiley-VCH Verlag; 2014. p. 1021–7.
62. Gabrilovich DI.
Myeloid-derived suppressor cells. Cancer Immunol Res. 2017 Jan 1;5(1):3–8.
63. Condamine T, Ramachandran I, Youn JI, Gabrilovich DI. Regulation of tumor metastasis by
myeloid-derived suppressor cells. Annu Rev Med. 2015 Jan 14;66:97–110.
64. Fujimura T, Kambayashi
Y, Fujisawa Y, Hidaka T, Aiba S. Tumor-associated
macrophages: Therapeutic targets for skin cancer. Vol. 8, Frontiers in
Oncology. Frontiers Media S.A.; 2018.
65. Jayasingam SD, Citartan M, Thang TH, Mat Zin AA, Ang KC, Ch’ng ES.
Evaluating the Polarization of Tumor-Associated Macrophages Into
M1 and M2 Phenotypes in Human Cancer Tissue: Technicalities and Challenges in
Routine Clinical Practice. Vol. 9, Frontiers in Oncology. Frontiers Media S.A.;
2020.
66. Nishimura H, Honjo T. PD-1: an inhibitory
immunoreceptor involved in peripheral tolerance. Trends Immunol. 2001
May;22(5):265–8.
67. Freeman GJ, Long AJ, Iwai Y, Bourque K,
Chernova T, Nishimura H, et al. Engagement of the PD-1 Immunoinhibitory
Receptor by a Novel B7 Family Member Leads to Negative Regulation of Lymphocyte
Activation [Internet]. Vol. 192, J. Exp. Med. 2000. Available from:
http://www.jem.org/cgi/content/full/192/7/1027
68. Zerdes I, Matikas A, Bergh J, Rassidakis
GZ, Foukakis T. Genetic, transcriptional and
post-translational regulation of the programmed death protein ligand 1 in
cancer: biology and clinical correlations. Vol. 37, Oncogene. Nature Publishing
Group; 2018. p. 4639–61.
69. Furue M, Ito T,
Wada N, Wada M, Kadono T, Uchi
H. Melanoma and Immune Checkpoint Inhibitors. Vol. 20, Current Oncology
Reports. Current Medicine Group LLC 1; 2018.
70. Sansom DM. CD28, CTLA-4 and their ligands:
Who does what and to whom? Vol. 101, Immunology. 2000. p. 169–77.
71. Trindade FM, de Freitas MLP, Bittencourt FV. Dermoscopic evaluation of superficial spreading melanoma. An Bras Dermatol. 2021 Mar 1;96(2):139–47.
72. Marghoob NG, Liopyris
K, Jaimes N. Dermoscopy: A review of the structures
that facilitate melanoma detection. Vol. 119, Journal of the American
Osteopathic Association. American Osteopathic Association; 2019. p. 380–90.
73. Dinnes J, Deeks JJ, Grainge MJ, Chuchu N, Ferrante di Ruffano L, Matin RN, et al. Visual inspection for
diagnosing cutaneous melanoma in adults. Vol. 2018, Cochrane Database of
Systematic Reviews. John Wiley and Sons Ltd; 2018.
74. Maron RC, Utikal
JS, Hekler A, Hauschild A, Sattler E, Sondermann W, et al. Artificial
intelligence and its effect on dermatologists’ accuracy in dermoscopic
melanoma image classification: Web-based survey study. J Med Internet Res. 2020
Sep 1;22(9).
75. Dinnes J, Deeks JJ, Chuchu N, Ferrante di Ruffano
L, Matin RN, Thomson DR, et al. Dermoscopy, with and
without visual inspection, for diagnosing melanoma in adults. Vol. 2018,
Cochrane Database of Systematic Reviews. John Wiley and Sons Ltd; 2018.
76. Davis LE, Shalin SC, Tackett AJ. Utility of
histone H3K27me3 and H4K20me as diagnostic indicators of melanoma. Melanoma
Res. 2020;159–65.
77. High WA. Detection of Genetic Aberrations in
the Assessment and Prognosis of Melanoma. Vol. 35, Dermatologic Clinics. W.B.
Saunders; 2017. p. 525–36.
78. Dinnes J, Di Ruffano LF, Takwoingi Y, Cheung
ST, Nathan P, Matin RN, et al. Ultrasound, CT, MRI, or PET-CT for staging and
re-staging of adults with cutaneous melanoma. Vol. 2019, Cochrane Database of
Systematic Reviews. John Wiley and Sons Ltd; 2019.
79. Testori AAE, Blankenstein SA, van Akkooi ACJ. Primary Melanoma: from History to Actual
Debates. Vol. 21, Current Oncology Reports. Springer; 2019.
80. Aderhold K, Wilson M, Berger AC, Levi S,
Bennett J. Precision Medicine in the Treatment of Melanoma. Vol. 29, Surgical
Oncology Clinics of North America. W.B. Saunders; 2020. p. 1–13.
81. Wellbrock C, Karasarides
M, Marais R. The RAF proteins take centre stage. Vol.
5, Nature Reviews Molecular Cell Biology. 2004. p. 875–85.
82. Pasquali S, Hadjinicolaou
A V., Chiarion Sileni V,
Rossi CR, Mocellin S. Systemic treatments for
metastatic cutaneous melanoma. Vol. 2018, Cochrane Database of Systematic
Reviews. John Wiley and Sons Ltd; 2018.
83. Leonardi GC, Falzone L, Salemi R, Zanghì A, Spandidos DA, Mccubrey
JA, et al. Cutaneous melanoma: From pathogenesis to therapy (Review). Vol. 52,
International Journal of Oncology. Spandidos
Publications; 2018. p. 1071–80.